CODA – Proteomics
Proteomics is an approach to catalogue and quantify the vast majority of proteins within any living cells and samples. As proteins are the workhorses of cellular biochemical reactions, identifying their identity and molecular abundance through this approach become an indispensable tool for understanding physiological functions in diverse organisms.
Extracted protein samples are often run through nano-Liquid Chromatography-tandem Mass Spectrometry (nanoLC-MS/MS) or matrix-assisted laser desorption/ionization- tandem mass spectrometry (MALDI-MS/MS) for protein and peptide mass profiling. However, researchers often encounter challenging downstream data analysis as the generated raw file formats can be diverse between service providers and MS instruments (either from Bruker, Thermo, Sciex, Agilent etc.) with a plethora of proteomics analysis software to choose from.
CODA-Proteomics undertakes this endeavour by streamlining proven and meticulous data analysis workflow for single protein identification as well as large-scale (shotgun) proteomics to cater raw file processing from various instrumentations and experimental complexity. As part of our commitment to data sharing and open science, proteomics datasets can now be uploaded to ProteomeXchange database to obtain unique accession numbers for publication purposes through our dedicated professional service.
PROTEOMICS WORKFLOW
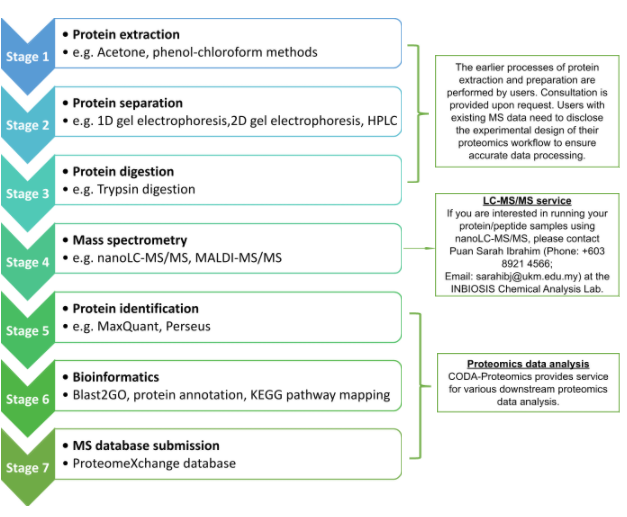
WHAT DO WE OFFER?
Overview of services
| Data analysis and bioinformatics | Software packages: MaxQuant, Perseus, Blast2Go, KOBAS, KEGG etc. depending on the users’ request and data complexity |
| Data sharing | Dataset deposition to ProteomeXchange database to obtain unique identifier for publication purposes |
PRICING DETAILS
| TYPE OF ANALYSIS | CHARGES |
| Single protein identification Data analysis of raw nano-Liquid Chromatography-Mass Spectrometry (nanoLC-MS) data from various systems (Thermo Orbitrap, Bruker, Agilent, Sciex etc.) | RM 100 / sample raw data* |
| Shotgun proteomics/large-scale protein identification Data analysis of raw nano-Liquid Chromatography-Mass Spectrometry (nanoLC-MS) data from various systems (Thermo Orbitrap, Bruker, Agilent, Sciex etc.) | RM 1000 / project (a total of 6 samples raw data**) |
| Data preparation and delivery to ProteomeXchange database Raw data storage and obtaining dataset identifier for publication | RM 25 / sample raw data* |
**If a project contains less or more than the 6 samples raw datasets, please discuss with CODA-Proteomics PIC for an updated charge.
# Analytical service (LC-MS/MS) is charged separately, subject to change according to customer needs.
ACHIEVEMENTS
PERSON IN CHARGE

University Lecturer DS13
hamizahshahirah@ukm.edu.my
Proteomics and Molecular Neuroscience

Science Officer C9 / Proteomics, Cell culture & Microscopy Lab PIC
+603 8921 4558
munirah@ukm.edu.my

